Highlight:
AnewOmni is the first all-atom geometric latent diffusion framework capable of designing peptides, antibodies, and small molecules within a unified generative model, demonstrating superior performance and cross-domain knowledge transfer in de novo binder design.
Overview
The design of target-specific binders—ranging from small molecules to peptides and antibodies—is fundamental to modern drug discovery. Historically, computational approaches have treated these molecular types as distinct domains, utilizing specialized generative models for each. This separated paradigm fails to leverage cross-domain transferability and limits the exploration of versatile therapeutic strategies. We introduce AnewOmni, a novel framework that unifies the design of diverse molecular binders into a single, cohesive model.
Methodology
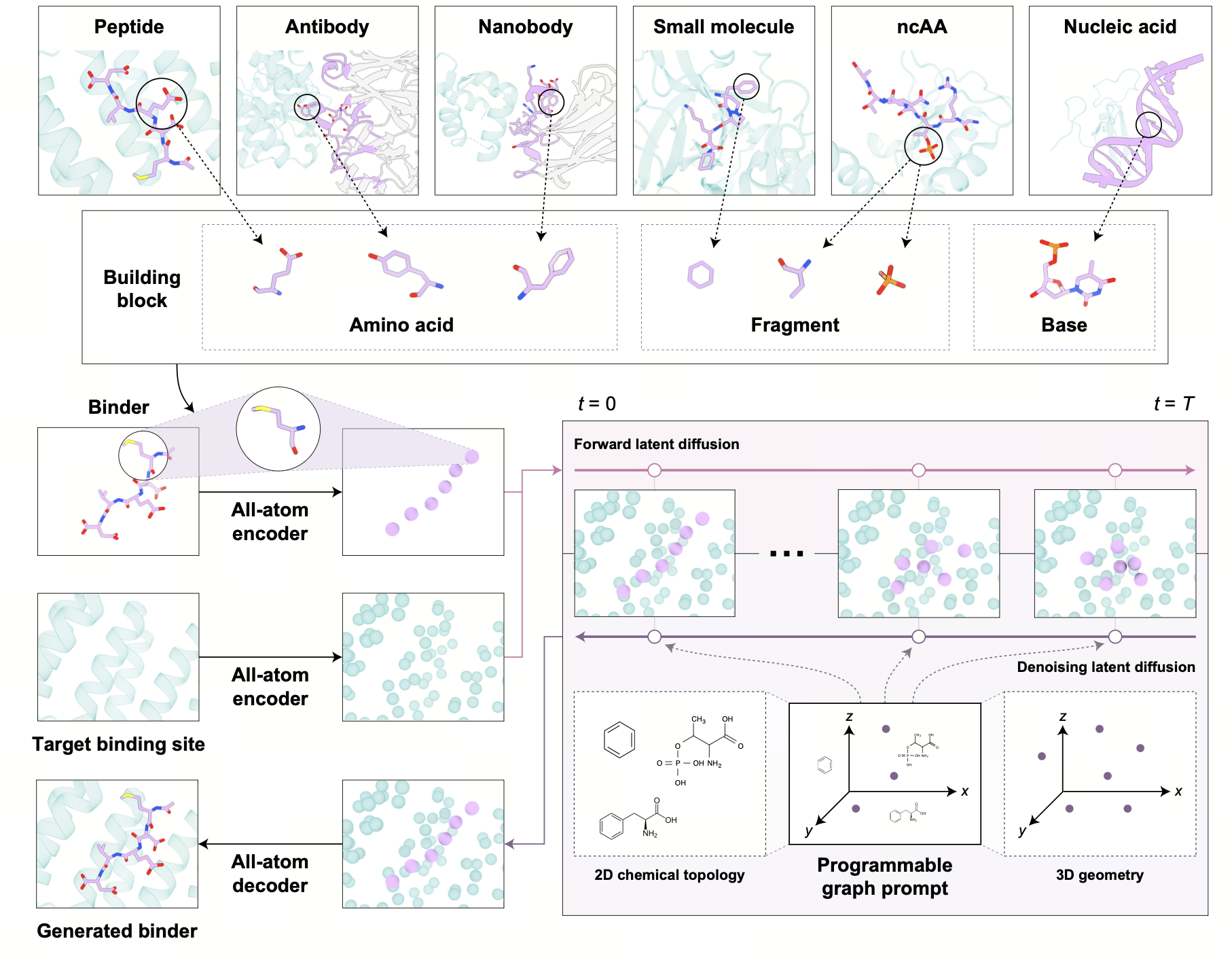
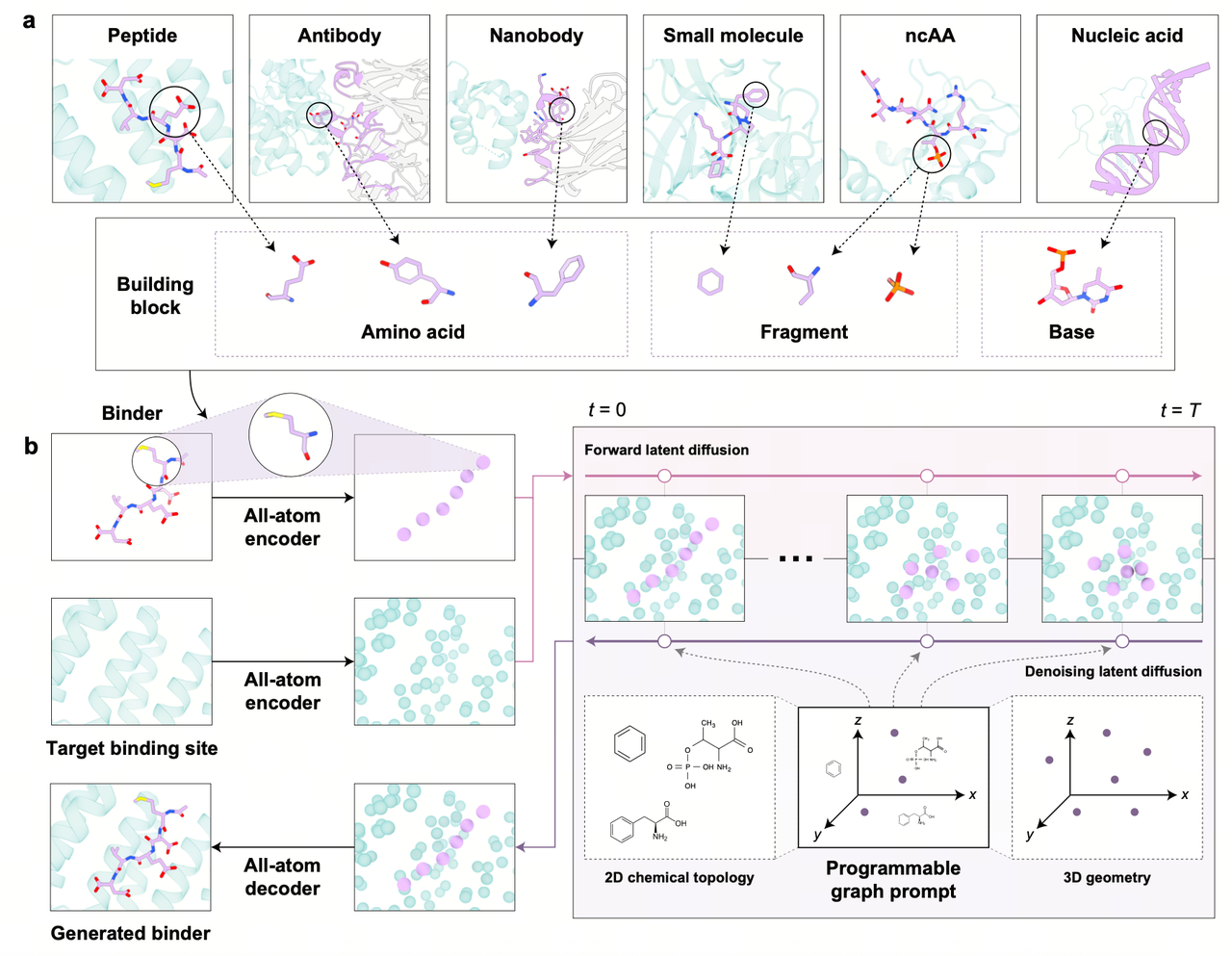
The core innovation of AnewOmni is its ability to process different biomolecular using a unified, full-atom representation called “blocks.” In this system, a block corresponds to either a standard amino acid (for peptides and antibodies) or a molecular fragment (for small molecules) identified via a principal subgraph algorithm. This representation preserves critical full-atom geometric details while capturing hierarchical structural priors.
To generate new binders, the framework employs a geometric latent diffusion model. The process begins with an iterative full-atom variational autoencoder (VAE), which compresses the “blocks” into a latent space defined by low-dimensional hidden states and spatial coordinates. An E(3)-equivariant diffusion model then operates within this compressed latent space to generate the binder’s structure. This latent-space approach allows the model to focus on global structural arrangements efficiently, while a specialized decoder reconstructs the fine-grained atomic details and chemical bonds.
Performance and Cross-Domain Learning
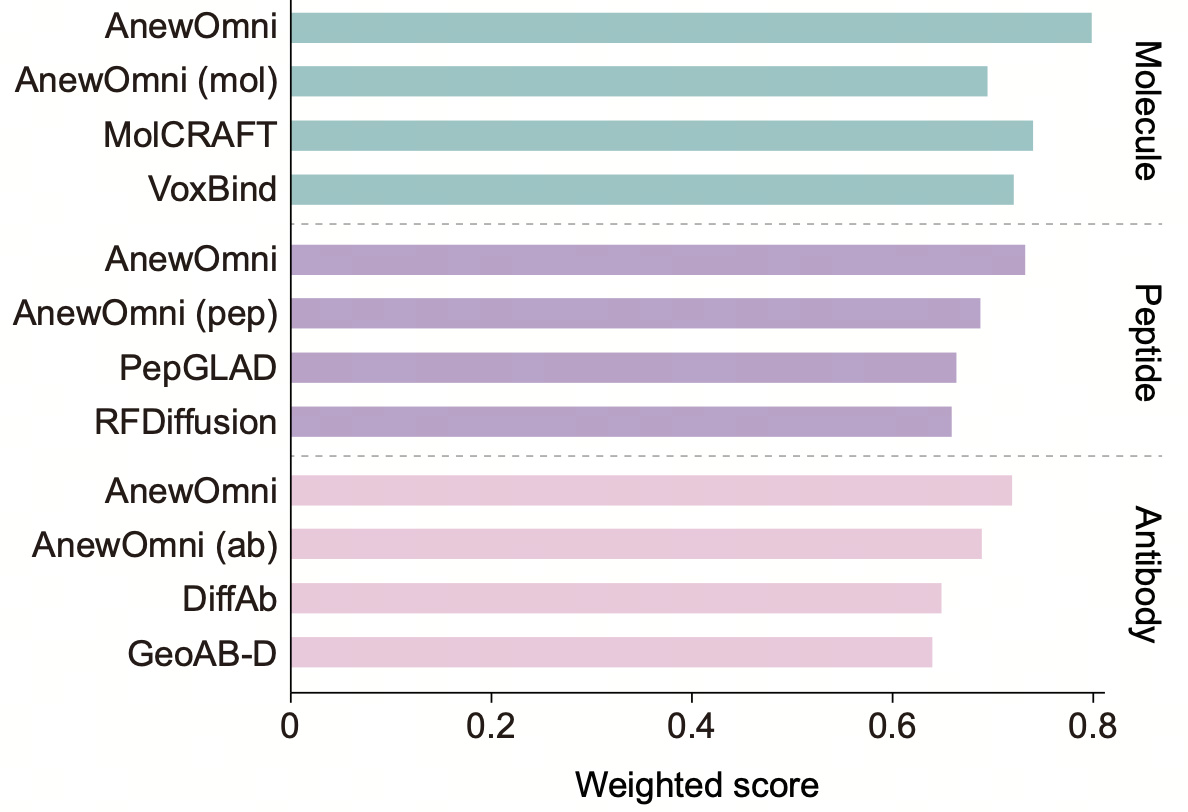
We benchmarked AnewOmni extensively against state-of-the-art, domain-specific models across peptide, antibody, and small molecule tasks. The unified framework consistently demonstrated superior performance in key metrics, including binding energy and structural validity.
AnewOmni leverages the inherent advantages of multi-domain training within a unified architecture, utilizing cross-domain knowledge transfer to refine all-atom interaction patterns and geometric precision across diverse molecular classes. This approach outperformed variants trained on single domains, indicating that AnewOmni successfully learns and transfers fundamental physical principles governing molecular interactions across different chemical modalities.
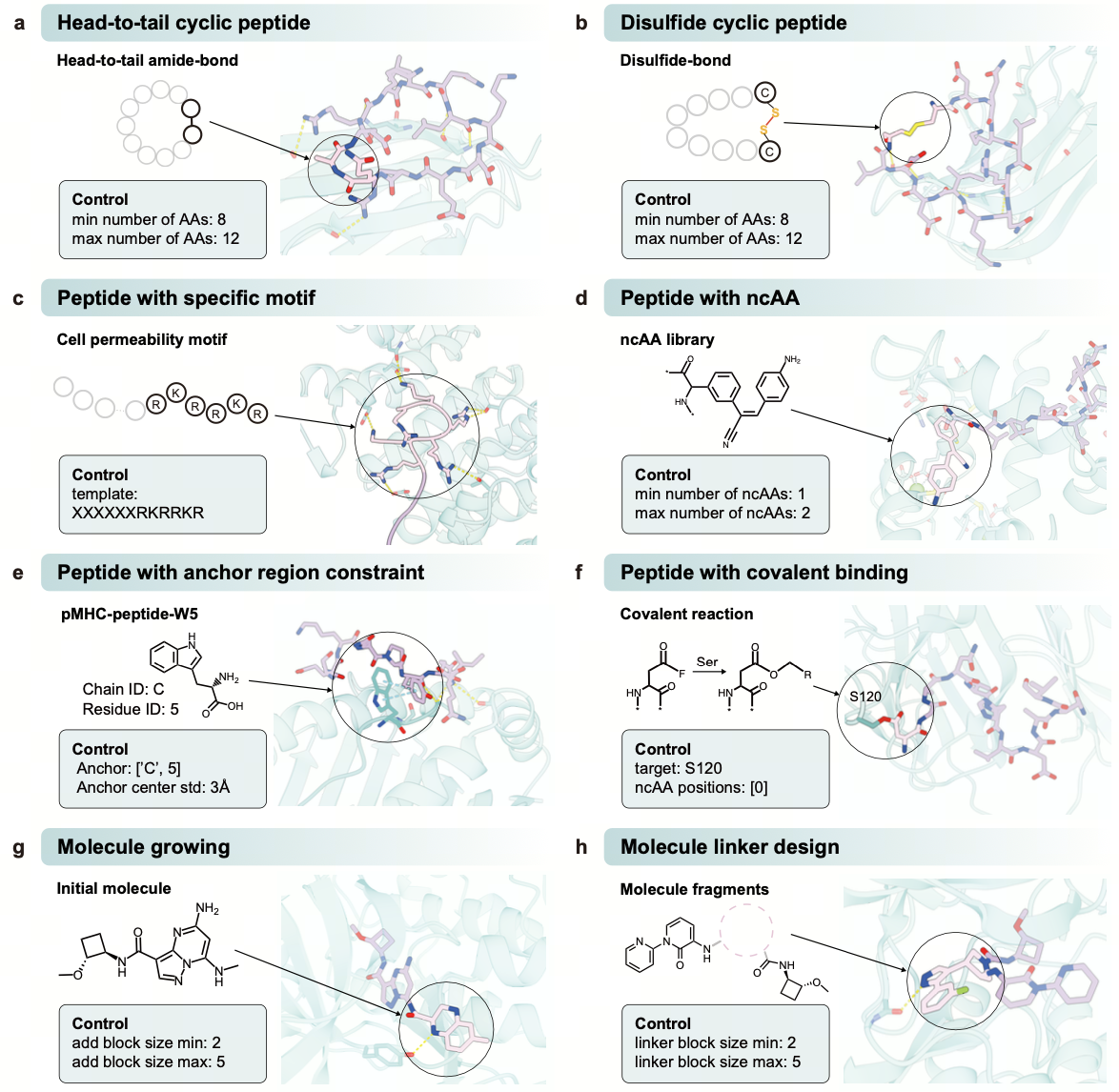
Additionally, AnewOmni exbihits highly programmability beyond standard de novo design. By flexibly manipulating 2D topology and 3D geometric “prompt”, the model can tackle specialized tasks such as small molecule fragment growing and linker design. This adaptability also extends to the generation of novel modalities—including cyclic peptides and peptides containing non-canonical amino acids—positioning the model as a versatile solution for a wide array of therapeutic biomolecular design scenarios.
Applications
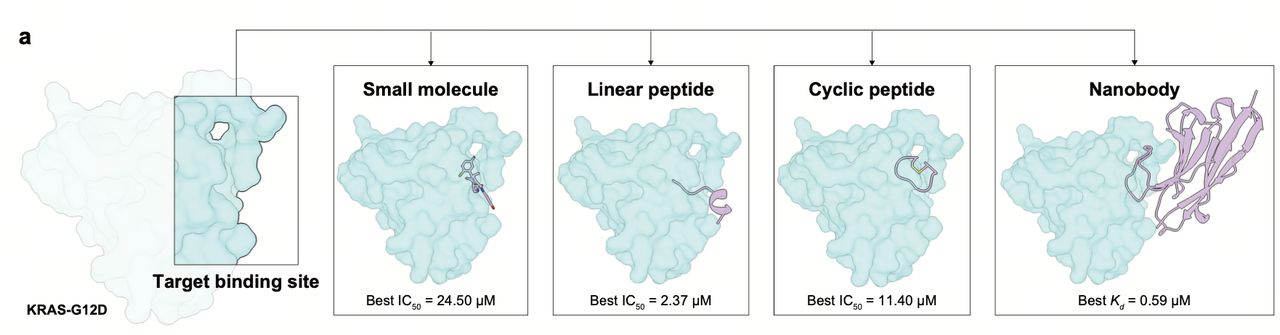
The versatility of AnewOmni is demonstrated through its ability to generate high-affinity binders across all three modalities for a single target. In a case study targeting KRAS G12D Switch II pocket, the model successfully designed small molecules, peptides, and nanobodies, achieving experimental success rates of 29% to 75%.
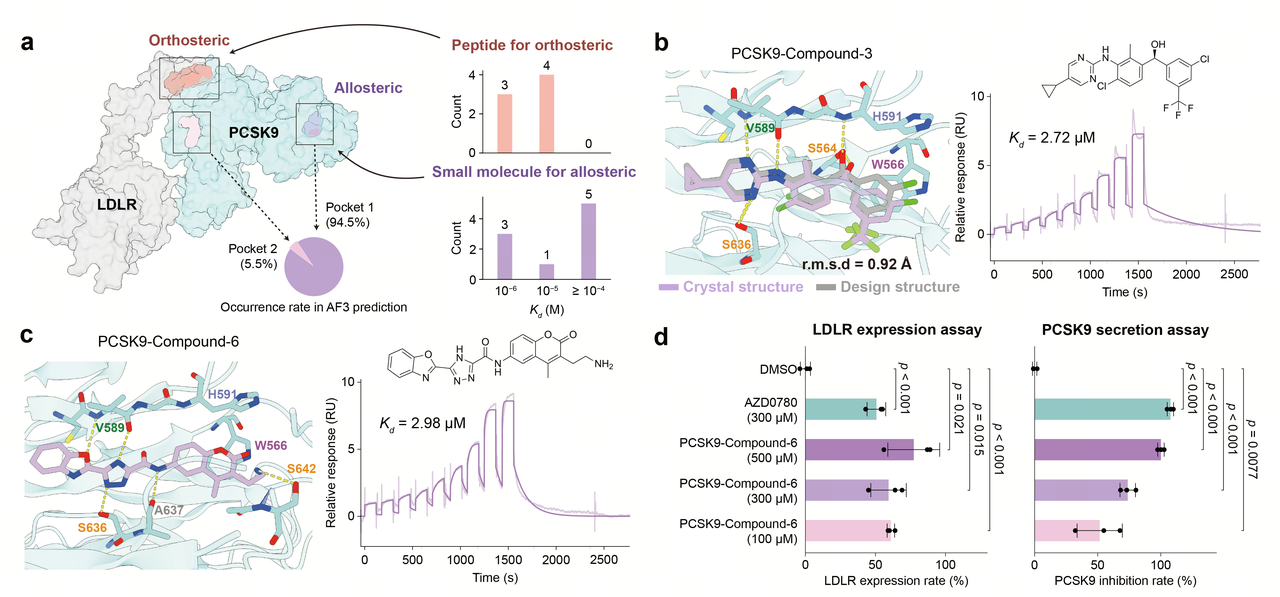
In a separate application targeting PCSK9, we leveraged AnewOmni to generate orthosteric peptides designed to disrupt its interaction with LDLR, as well as to discover small molecules targeting a newly identified allosteric site. The generated candidates exhibited strong binding affinities, which were confirmed by Surface Plasmon Resonance (SPR) measurements. We further characterized PCSK9-compound-3, a small-molecule inhibitor demonstrating potent binding affinity, using X-ray crystallography. This structural analysis revealed high fidelity, with the experimentally resolved structure closely matching the model-predicted binding pose (RMSD 0.92 Å). Another highly potent candidate is PCSK9-compound-6 which demonstrated a binding affinity (Kd) of 2.98 µM. To evaluate its functional impact, we advanced this compound into cellular assays measuring LDLR expression. Remarkably, treatment with PCSK9-compound-6 at 100 µM upregulated LDLR expression to levels comparable to those achieved by the clinical-stage reference compound, AZD0780, at a much higher concentration of 300 µM. Furthermore, cellular assays confirmed that PCSK9-compound-6 effectively suppresses PCSK9 secretion.
By bridging the gap between distinct chemical spaces, AnewOmni serves as a powerful tool for holistic drug discovery, enabling the simultaneous exploration of multiple therapeutic modalities.
